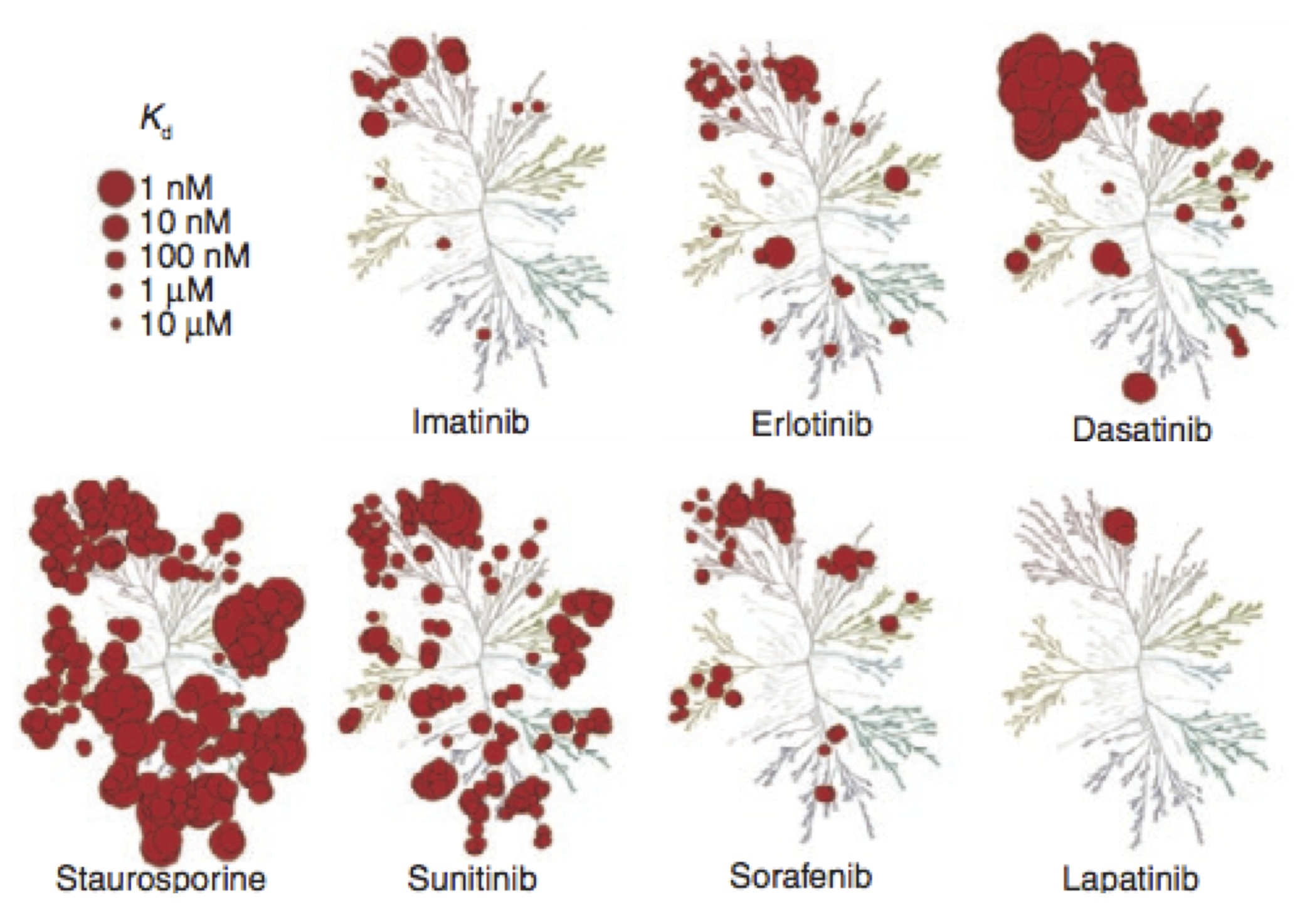
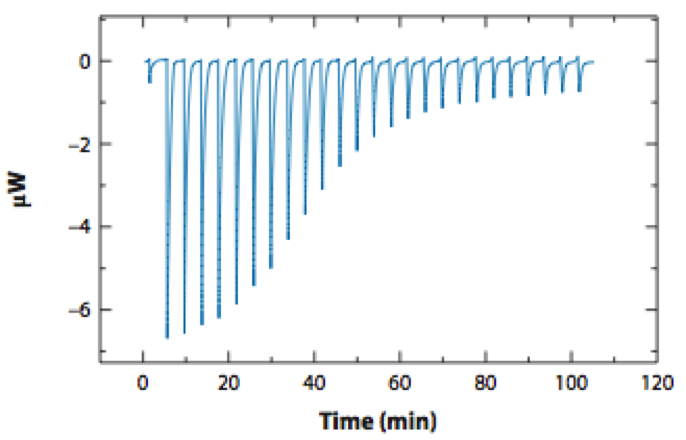
The Chodera lab uses computation and experiment to develop quantitative, multiscale models of the effects of small molecules on biomolecular macromolecules and cellular pathways and understand the functional and therapeutic ramifications of mutations. The group utilizes physical models, rigorous statistical mechanics, and open source software development practices with overall goals of engineering novel therapeutics and tools for chemical biology, predicting resistance or susceptibility to therapy, and understanding the physical driving forces behind the emergence of drug resistance. We develop and use advanced algorithms for molecular dynamics simulations on GPUs and distributed computing platforms, in addition to high-throughput experiments to characterize biophysical interactions between small molecules and their targets.
Our lab is a core member of the AI-driven Structure-enabled Antiviral Discovery Platform (ASAP), the Folding@home Consortium, the Open Force Field Initiative, and the COVID Moonshot.
FEATURED RESEARCH PROJECTS