SEARCH
ARTICLE TYPES
Spectral rate theory for two-state kinetics
/Jan-Hendrik Prinz, John D. Chodera, and Frank Noé.
Phys. Rev. X 4:011020, 2014. [DOI] [PDF]
We present a new mathematical framework for unifying various two-state rate theories presented in the physical chemistry literature over many decades, and provide a quantitative way to measure reaction coordinate quality.
Markov models of molecular kinetics: Generation and validation
/Jan-Hendrik Prinz, Hao Wu, Marco Sarich, Bettina Keller, Martin Fischbach, Martin Held, John D. Chodera, Christof Schüttle, and Frank Noé.
J. Chem. Phys. 134:174105, 2011. [DOI] [PDF]
A review of current best practices for the generation and validation of Markov state models for describing the stochastic dynamics of biomolecular systems.
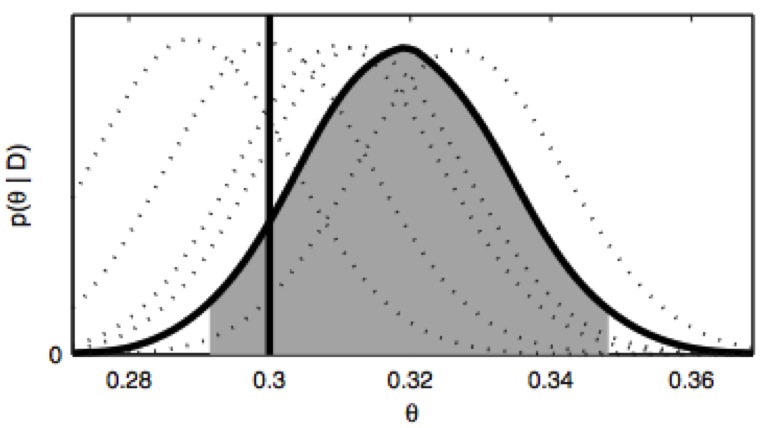
Bayesian comparison of Markov models of molecular dynamics with detailed balance constraint
/Sergio Bacallado, John D. Chodera, and Vijay S. Pande.
J. Chem. Phys. 131:045106, 2009. [DOI] [PDF]
A Bayesian scheme for comparing state space decompositions for Markov state models of biomolecular dynamics that incorporates the fact that physical systems must obey detailed balance. This paper utilizes recent results from Markov chain theory on edge-reinforced random walks.
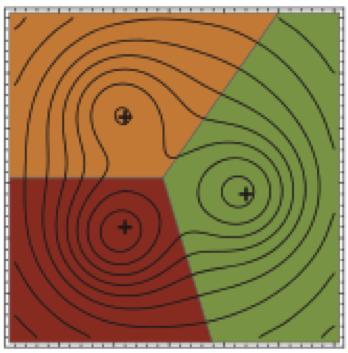
Automatic discovery of metastable states for the construction of Markov models of macromolecular conformational dynamics
/John D. Chodera*, Nina Singhal*, William C. Swope, Jed W. Pitera, Vijay S. Pande, and Ken A. Dill.
J. Chem. Phys. 126:155101, 2007. [DOI] [PDF]
Proposing one of the first automated algorithms for discovering kinetically metastable states of biomolecules from molecular simulations, this paper shows how many biomolecules can possess numerous distinct long-lived conformational states even though the the equilibrium populations of these states may too small for standard structural biology techniques to detect.
Long-time protein folding dynamics from short-time molecular dynamics simulations
/John D. Chodera, William C. Swope, Jed W. Pitera, and Ken A. Dill.
Multiscale Model. Simul. 5:1214, 2006. [DOI] [PDF]
We show how the long-time dynamics of biomolecular systems can be recapitulated from statistics collected from short molecular simulations sampling transitions between kinetically metastable states.