Escaping atom types in force fields using direct chemical perception
/David L. Mobley, Caitlin C. Bannan, Andrea Rizzi, Christopher I. Bayly, John D. Chodera, Victoria T Lim, Nathan M. Lim, Kyle A. Beauchamp, Michael R. Shirts, Michael K. Gilson, Peter K. Eastman.
Journal of Chemical Theory and Computation 14:6076, 2018 [DOI] [bioRxiv]
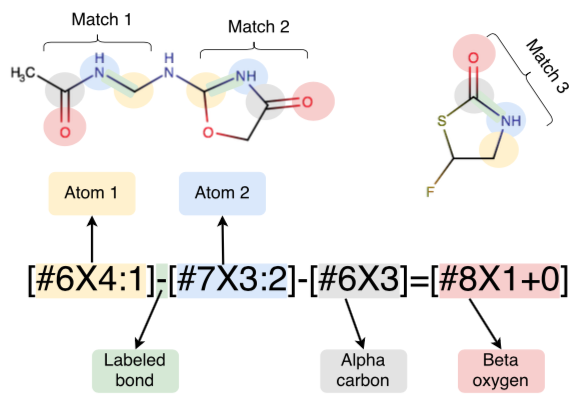
We describe the philosophy behind a modern approach to molecular mechanics forcefield parameterization, and present initial results for the first SMIRNOFF-encoded forcefield: SMIRNOFF99Frosst.