Quantifying configuration-sampling error in Langevin simulations of complex molecular systems
/Josh Fass, David Sivak , Gavin E. Crooks, Kyle A. Beauchamp, Benedict Leimkuhler, and John Chodera.
Entropy 20:318, 2018. [DOI] [PDF] [GitHub] [bioRxiv preprint]
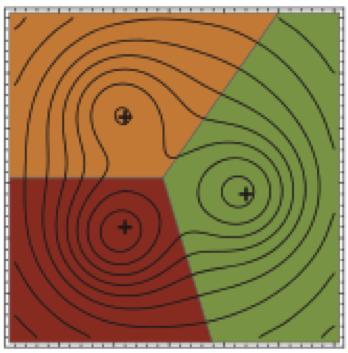
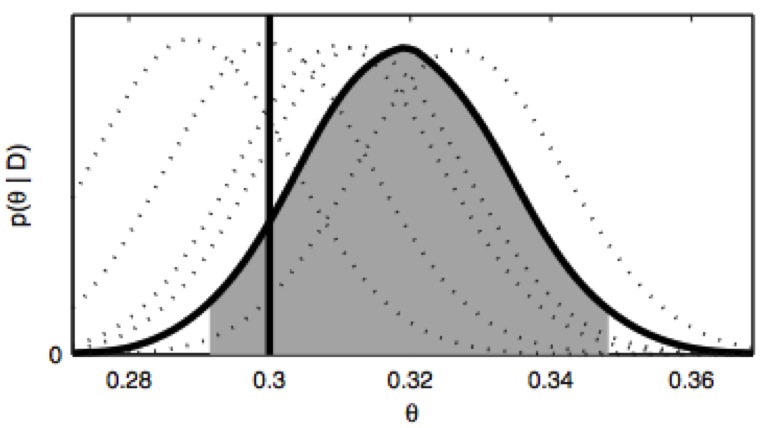
Molecular dynamics simulations necessarily use a finite timestep, which introduces error or bias in the sampled configuration space density that grows rapidly with increasing timestep. For the first time, we show how to compute a natural measure of this error---the KL divergence---in both phase and configuration space for a widely used family of Langevin integrators, and show that VRORV is generally superior for simulation of molecular systems.