The limitations of constant-force-feedback experiments
/Phillip J. Elms, John D. Chodera, Carlos J. Bustamante, and Susan Marqusee.
Biophys. J. 103:1490, 2012. [DOI] [PDF]
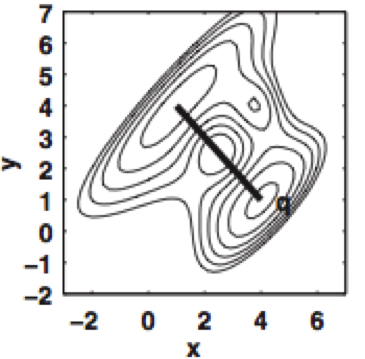
Popular constant-force-feedback single-molecule experiments can cause severe artifacts in single-molecule force spectroscopy data. We demonstrate a simple alternative that eliminates these artifacts.