SEARCH
ARTICLE TYPES
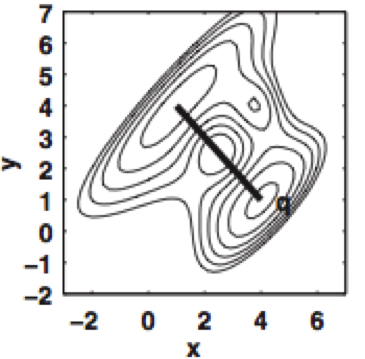
Spectral rate theory for two-state kinetics
/Jan-Hendrik Prinz, John D. Chodera, and Frank Noé.
Phys. Rev. X 4:011020, 2014. [DOI] [PDF]
We present a new mathematical framework for unifying various two-state rate theories presented in the physical chemistry literature over many decades, and provide a quantitative way to measure reaction coordinate quality.
Time step rescaling recovers continuous-time dynamical properties for discrete-time Langevin integration of nonequilibrium systems
/David A. Sivak, John D. Chodera, and Gavin E. Crooks.
J. Phys. Chem. B, 118:6466-6474, 2014. William C. Swope Festschrift issue. [DOI] [PDF]
We derive a simple, easy-to-implement Langevin integrator that has universally useful properties in molecular simulations.
Keywords: velocity Verlet with velocity randomization; VVVR; nonequilibrium integration
Identifying ligand binding sites and poses using GPU-accelerated Hamiltonian replica exchange molecular dynamics
/Kai Wang K, John D. Chodera, Yanzhi Yang, and Michael R. Shirts.
J. Comput. Aid. Mol. Des. 27:989, 2013. [DOI] [PDF]
We show how bound ligand poses can be identified even when the location of the binding sites are unknown using the machinery of alchemical modern free energy calculations on graphics processors.
Systematic improvement of a classical molecular model of water
/Lee-Ping Wang, Teresa L. Head-Gordon, Jay W. Ponder, Pengyu Ren, John D. Chodera, Peter K. Eastman, Todd J. Martinez, and Vijay S. Pande.
J. Phys. Chem. B 117:9956, 2013. [DOI] [PDF]
A new inexpensive polarizable model of liquid water for next-generation forcefields is derived using an automated parameterization engine.
Using nonequilibrium fluctuation theorems to understand and correct errors in equilibrium and nonequilibrium discrete Langevin dynamics simulations
/David A. Sivak, John D. Chodera, and Gavin E. Crooks.
Phys. Rev. X 3:011007, 2013. [DOI] [PDF]
The finite-timestep errors in molecular dynamics simulations can be interpreted as a form of nonequilibrium work. We show how this leads to straightforward schemes for correcting for these errors or assessing their impact.
Keywords: velocity verlet with Velocity randomization; VVVR; nonequilibrium free energy; integrator error; nonequilibrium integration
OpenMM 4: A reusable, extensible, hardware independent library for high performance molecular simulation
/Peter Eastman, Mark S. Friedrichs, John D. Chodera, Randy J. Radmer, Chris M. Bruns, Joy P. Ku, Kyle A. Beauchamp, T. J. Lane, Lee-Ping Wang, Diwakar Shukla, Tony Tye, Mike Houston, Timo Stich, Christoph Klein, Michael R. Shirts, and Vijay S. Pande.
J. Chem. Theor. Comput. 9:461, 2013. [DOI] [PDF]
We describe the latest version of an open-source, GPU-accelerated library and toolkit for molecular simulation.
The limitations of constant-force-feedback experiments
/Phillip J. Elms, John D. Chodera, Carlos J. Bustamante, and Susan Marqusee.
Biophys. J. 103:1490, 2012. [DOI] [PDF]
Popular constant-force-feedback single-molecule experiments can cause severe artifacts in single-molecule force spectroscopy data. We demonstrate a simple alternative that eliminates these artifacts.
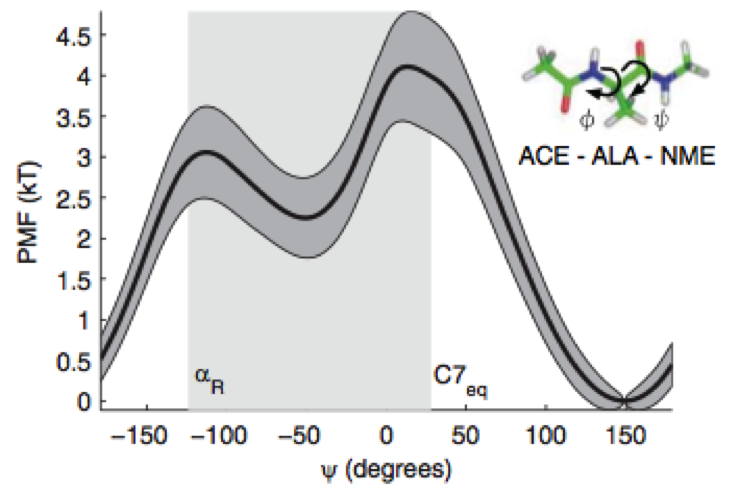
On the use of experimental observations to bias simulated ensembles
/Jed W. Pitera and John D. Chodera.
J. Chem. Theor. Comput. 8:3445, 2012. [DOI] [PDF]
We show how the concept of maximum entropy can be used to recover unbiased conformational distributions from experimental data, and how this concept relates to the popular `ensemble refinement' schemes for NMR data analysis.
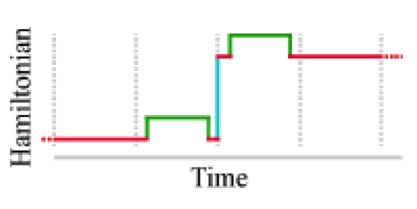
Nonequilibrium candidate Monte Carlo is an efficient tool for equilibrium simulation
/Jerome P. Nilmeier, Gavin E. Crooks, David D. L. Minh, and John D. Chodera.
Proc. Natl. Acad. Sci. USA 108:E1009, 2011. [DOI] [PDF]
We present a significant generalization of Monte Carlo methods that provide an enormously useful tool for enhancing the efficiency of molecular simulations and enabling molecular design.
Keywords: NCMC; Monte Carlo; Metropolis-Hastings; acceptance rates; molecular dynamics
A robust approach to estimating rates from time-correlation functions
/John D. Chodera, Phillip J. Elms, William C. Swope, Jan-Hendrik Prinz, Susan Marqusee, Carlos Bustamante, Frank Noé, Vijay S. Pande
Preprint ahead of submission: [arXiv] [PDF] [SI]
The estimation of rates from experimental single-molecule data is fraught with peril. We describe some of the failures of existing methods and suggest a robust way to estimate rates from time-correlation functions.
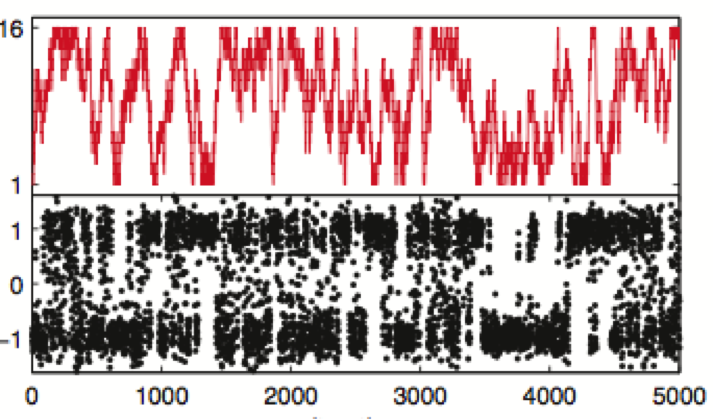
Bayesian hidden Markov model analysis of single-molecule force spectroscopy: Characterizing kinetics under measurement uncertainty
/John D. Chodera, Phillip Elms, Frank Noé, Bettina Keller, Christian M. Kaiser, Aaron Ewall-Wice, Susan Marqusee, Carlos Bustamante, and Nina Singhal Hinrichs.
preprint: [arXiv]
We describe the general theory and implementation for a Bayesian extension of hidden Markov models applicable to the characterization of how measurement uncertainty and finite statistics can impact the confidence in rate constants and conformational state properties.
Dynamical reweighting: Improved estimates for dynamical properties from simulations at multiple temperatures
/John D. Chodera, William C. Swope, Frank Noé, Jan-Hendrik Prinz, Michael R. Shirts, and Vijay S. Pande.
J. Chem. Phys. 134:244107, 2011. [DOI] [PDF]
We describe how reweighing techniques can provide optimal estimates of temperature-dependent dynamical properties from simulations conducted at multiple temperatures.